Spatial simultaneous autoregressive SAC model estimation by GMM
gstsls.RdAn implementation of Kelejian and Prucha's generalised moments estimator for the autoregressive parameter in a spatial model with a spatially lagged dependent variable.
Usage
gstsls(formula, data = list(), listw, listw2 = NULL, na.action = na.fail,
zero.policy = attr(listw, "zero.policy"), pars=NULL, scaleU=FALSE, control = list(),
verbose=NULL, method="nlminb", robust=FALSE, legacy=FALSE, W2X=TRUE, sig2n_k=FALSE)
# S3 method for class 'Gmsar'
impacts(obj, ..., n = NULL, tr = NULL, R = NULL,
listw = NULL, evalues=NULL, Q=NULL)Arguments
- formula
a symbolic description of the model to be fit. The details of model specification are given for
lm()- data
an optional data frame containing the variables in the model. By default the variables are taken from the environment which the function is called.
- listw
a
listwobject created for example bynb2listw- listw2
a
listwobject created for example bynb2listw, if not given, set to the same spatial weights as the listw argument- na.action
a function (default
na.fail), can also bena.omitorna.excludewith consequences for residuals and fitted values - in these cases the weights list will be subsetted to remove NAs in the data. It may be necessary to set zero.policy to TRUE because this subsetting may create no-neighbour observations. Note that only weights lists created without using the glist argument tonb2listwmay be subsetted.- zero.policy
default NULL, use global option value; if TRUE assign zero to the lagged value of zones without neighbours, if FALSE (default) assign NA - causing
GMerrorsar()to terminate with an error- pars
starting values for \(\lambda\) and \(\sigma^2\) for GMM optimisation, if missing (default), approximated from initial 2sls model as the autocorrelation coefficient corrected for weights style and model sigma squared
- scaleU
Default FALSE: scale the OLS residuals before computing the moment matrices; only used if the
parsargument is missing- control
A list of control parameters. See details in optim or nlminb
- verbose
default NULL, use global option value; if TRUE, reports function values during optimization.
- method
default nlminb, or optionally a method passed to optim to use an alternative optimizer
- robust
see
stsls- legacy
see
stsls- W2X
see
stsls- sig2n_k
see
stsls- obj
A spatial regression object created by
lagsarlm,lagmessor bylmSLX; inHPDinterval.LagImpact, a LagImpact object- ...
Arguments passed through to methods in the coda package
- tr
A vector of traces of powers of the spatial weights matrix created using
trW, for approximate impact measures; if not given,listwmust be given for exact measures (for small to moderate spatial weights matrices); the traces must be for the same spatial weights as were used in fitting the spatial regression, and must be row-standardised- evalues
vector of eigenvalues of spatial weights matrix for impacts calculations
- R
If given, simulations are used to compute distributions for the impact measures, returned as
mcmcobjects; the objects are used for convenience but are not output by an MCMC process- Q
default NULL, else an integer number of cumulative power series impacts to calculate if
tris given- n
defaults to
length(obj$residuals); in the method forGmsarobjects it may be used in panel settings to compute the impacts for cross-sectional weights only, suggested by Angela Parenti
Details
When the control list is set with care, the function will converge to values close to the ML estimator without requiring computation of the Jacobian, the most resource-intensive part of ML estimation.
Value
A list object of class Gmsar
- lambda
simultaneous autoregressive error coefficient
- coefficients
GMM coefficient estimates (including the spatial autocorrelation coefficient)
- rest.se
GMM coefficient standard errors
- s2
GMM residual variance
- SSE
sum of squared GMM errors
- parameters
number of parameters estimated
- lm.model
NULL
- call
the call used to create this object
- residuals
GMM residuals
- lm.target
NULL
- fitted.values
Difference between residuals and response variable
- formula
model formula
- aliased
NULL
- zero.policy
zero.policy for this model
- LL
NULL
- vv
list of internal bigG and litg components for testing optimisation surface
- optres
object returned by optimizer
- pars
start parameter values for optimisation
- Hcov
NULL
- na.action
(possibly) named vector of excluded or omitted observations if non-default na.action argument used
References
Kelejian, H. H., and Prucha, I. R., 1999. A Generalized Moments Estimator for the Autoregressive Parameter in a Spatial Model. International Economic Review, 40, pp. 509–533; Cressie, N. A. C. 1993 Statistics for spatial data, Wiley, New York.
Roger Bivand, Gianfranco Piras (2015). Comparing Implementations of Estimation Methods for Spatial Econometrics. Journal of Statistical Software, 63(18), 1-36. doi:10.18637/jss.v063.i18 .
Examples
#require("spdep", quietly=TRUE)
data(oldcol, package="spdep")
COL.errW.GM <- gstsls(CRIME ~ INC + HOVAL, data=COL.OLD, spdep::nb2listw(COL.nb, style="W"))
summary(COL.errW.GM)
#>
#> Call:
#> gstsls(formula = CRIME ~ INC + HOVAL, data = COL.OLD, listw = spdep::nb2listw(COL.nb,
#> style = "W"))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -37.87710 -5.74981 -0.21447 6.19490 22.85527
#>
#> Type: GM SARAR estimator
#> Coefficients: (GM standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> Rho_Wy 0.454171 0.177939 2.5524 0.010699
#> (Intercept) 43.782818 10.467857 4.1826 2.882e-05
#> INC -0.994831 0.366818 -2.7121 0.006687
#> HOVAL -0.267076 0.088149 -3.0298 0.002447
#>
#> Lambda: 0.016647
#> Residual variance (sigma squared): 94.921, (sigma: 9.7427)
#> GM argmin sigma squared: 93.729
#> Number of observations: 49
#> Number of parameters estimated: 6
#>
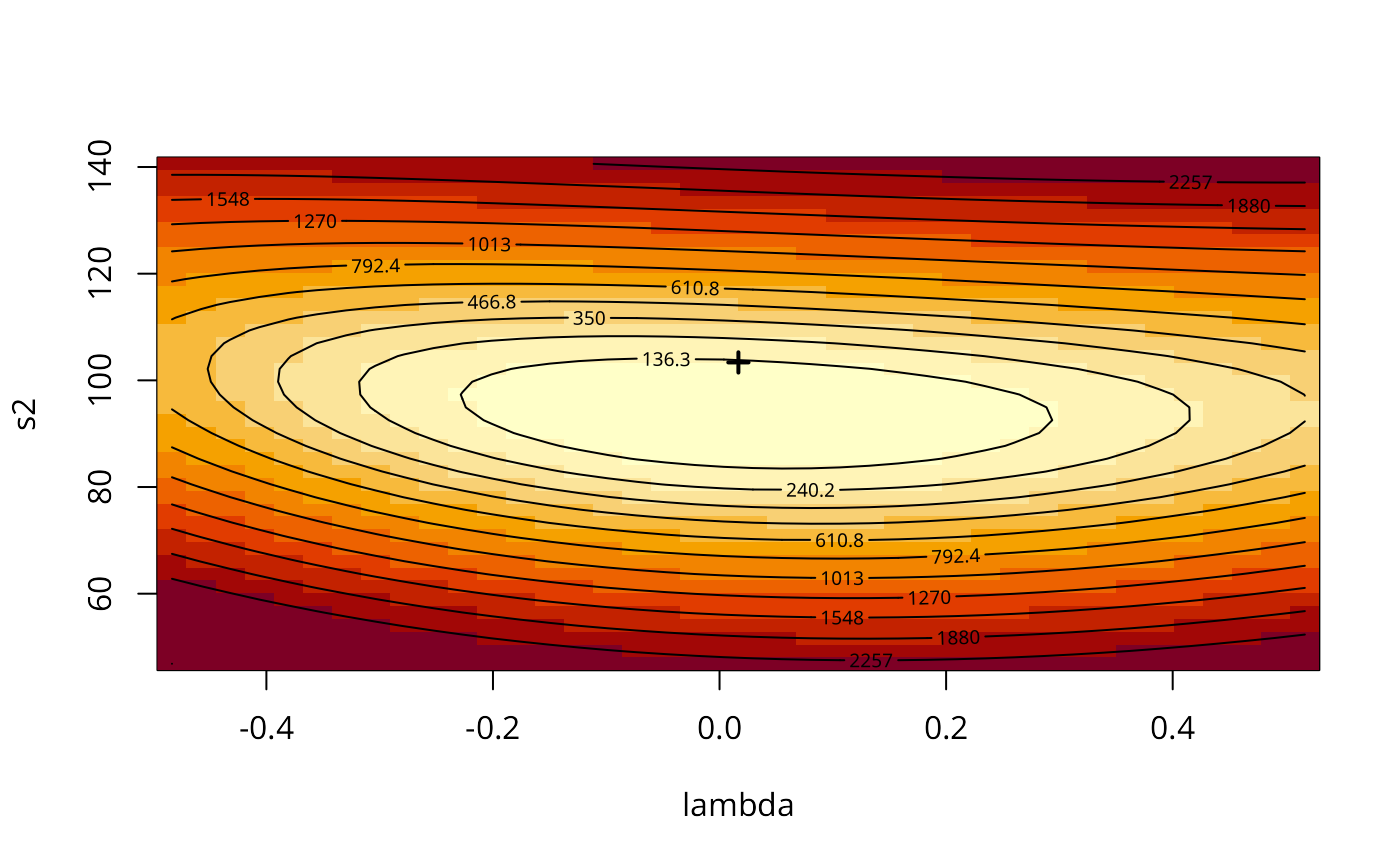
aa <- GMargminImage(COL.errW.GM)
levs <- quantile(aa$z, seq(0, 1, 1/12))
image(aa, breaks=levs, xlab="lambda", ylab="s2")
points(COL.errW.GM$lambda, COL.errW.GM$s2, pch=3, lwd=2)
contour(aa, levels=signif(levs, 4), add=TRUE)
 COL.errW.GM <- gstsls(CRIME ~ INC + HOVAL, data=COL.OLD,
spdep::nb2listw(COL.nb, style="W"), scaleU=TRUE)
summary(COL.errW.GM)
#>
#> Call:
#> gstsls(formula = CRIME ~ INC + HOVAL, data = COL.OLD, listw = spdep::nb2listw(COL.nb,
#> style = "W"), scaleU = TRUE)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -37.87710 -5.74981 -0.21447 6.19490 22.85527
#>
#> Type: GM SARAR estimator
#> Coefficients: (GM standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> Rho_Wy 0.454171 0.177939 2.5524 0.010699
#> (Intercept) 43.782818 10.467857 4.1826 2.882e-05
#> INC -0.994831 0.366818 -2.7121 0.006687
#> HOVAL -0.267076 0.088149 -3.0298 0.002447
#>
#> Lambda: 0.016647
#> Residual variance (sigma squared): 94.921, (sigma: 9.7427)
#> (scaled) GM argmin sigma squared: 0.96653
#> Number of observations: 49
#> Number of parameters estimated: 6
#>
listw <- spdep::nb2listw(COL.nb)
W <- as(listw, "CsparseMatrix")
trMat <- trW(W, type="mult")
impacts(COL.errW.GM, tr=trMat)
#> Impact measures (lag, trace):
#> Direct Indirect Total
#> INC dy/dx -1.0543769 -0.7682309 -1.8226078
#> HOVAL dy/dx -0.2830618 -0.2062420 -0.4893038
COL.errW.GM <- gstsls(CRIME ~ INC + HOVAL, data=COL.OLD,
spdep::nb2listw(COL.nb, style="W"), scaleU=TRUE)
summary(COL.errW.GM)
#>
#> Call:
#> gstsls(formula = CRIME ~ INC + HOVAL, data = COL.OLD, listw = spdep::nb2listw(COL.nb,
#> style = "W"), scaleU = TRUE)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -37.87710 -5.74981 -0.21447 6.19490 22.85527
#>
#> Type: GM SARAR estimator
#> Coefficients: (GM standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> Rho_Wy 0.454171 0.177939 2.5524 0.010699
#> (Intercept) 43.782818 10.467857 4.1826 2.882e-05
#> INC -0.994831 0.366818 -2.7121 0.006687
#> HOVAL -0.267076 0.088149 -3.0298 0.002447
#>
#> Lambda: 0.016647
#> Residual variance (sigma squared): 94.921, (sigma: 9.7427)
#> (scaled) GM argmin sigma squared: 0.96653
#> Number of observations: 49
#> Number of parameters estimated: 6
#>
listw <- spdep::nb2listw(COL.nb)
W <- as(listw, "CsparseMatrix")
trMat <- trW(W, type="mult")
impacts(COL.errW.GM, tr=trMat)
#> Impact measures (lag, trace):
#> Direct Indirect Total
#> INC dy/dx -1.0543769 -0.7682309 -1.8226078
#> HOVAL dy/dx -0.2830618 -0.2062420 -0.4893038