Spatial conditional and simultaneous autoregression model estimation
spautolm.RdFunction taking family and weights arguments for spatial autoregression model estimation by Maximum Likelihood, using dense matrix methods, not suited to large data sets with thousands of observations. With one of the sparse matrix methods, larger numbers of observations can be handled, but the interval= argument should be set. The implementation is GLS using the single spatial coefficient value, here termed lambda, found by line search using optimize to maximise the log likelihood.
Usage
spautolm(formula, data = list(), listw, weights,
na.action, family = "SAR", method="eigen", verbose = NULL, trs=NULL,
interval=NULL, zero.policy = NULL, tol.solve=.Machine$double.eps,
llprof=NULL, control=list())
# S3 method for class 'Spautolm'
summary(object, correlation = FALSE, adj.se=FALSE,
Nagelkerke=FALSE, ...)Arguments
- formula
a symbolic description of the model to be fit. The details of model specification are given for
lm()- data
an optional data frame containing the variables in the model. By default the variables are taken from the environment which the function is called.
- listw
a
listwobject created for example bynb2listw- weights
an optional vector of weights to be used in the fitting process
- na.action
a function (default
options("na.action")), can also bena.omitorna.excludewith consequences for residuals and fitted values - in these cases the weights list will be subsetted to remove NAs in the data. Note that only weights lists created without using the glist argument tonb2listwmay be subsetted.- family
character string: either
"SAR"or"CAR"for simultaneous or conditional autoregressions;"SMA"for spatial moving average added thanks to Jielai Ma -"SMA"is only implemented for method="eigen"because it necessarily involves dense matrices- method
character string: default
"eigen"for use of dense matrices,"Matrix_J"for sparse matrices (restricted to spatial weights symmetric or similar to symmetric) using methods in the Matrix package; “Matrix” provides updating Cholesky decomposition methods. Values of method may also include "LU", which provides an alternative sparse matrix decomposition approach, and the "Chebyshev" and Monte Carlo "MC" approximate log-determinant methods.- verbose
default NULL, use global option value; if TRUE, reports function values during optimization.
- trs
default NULL, if given, a vector of powered spatial weights matrix traces output by
trW; when given, used in some Jacobian methods- interval
search interval for autoregressive parameter when not using method="eigen"; default is c(-1,0.999),
optimizewill reset NA/NaN to a bound and gives a warning when the interval is poorly set; method="Matrix" will attempt to search for an appropriate interval, if find_interval=TRUE (fails on some platforms)- zero.policy
default NULL, use global option value; Include list of no-neighbour observations in output if TRUE — otherwise zero.policy is handled within the listw argument
- tol.solve
the tolerance for detecting linear dependencies in the columns of matrices to be inverted - passed to
solve()(default=double precision machine tolerance). Errors insolve()may constitute indications of poorly scaled variables: if the variables have scales differing much from the autoregressive coefficient, the values in this matrix may be very different in scale, and inverting such a matrix is analytically possible by definition, but numerically unstable; rescaling the RHS variables alleviates this better than setting tol.solve to a very small value- llprof
default NULL, can either be an integer, to divide the feasible range into llprof points, or a sequence of spatial coefficient values, at which to evaluate the likelihood function
- control
list of extra control arguments - see section below
- object
Spautolmobject fromspautolm- correlation
logical; if 'TRUE', the correlation matrix of the estimated parameters is returned and printed (default=FALSE)
- adj.se
if TRUE, adjust the coefficient standard errors for the number of fitted coefficients
- Nagelkerke
if TRUE, the Nagelkerke pseudo R-squared is reported
- ...
further arguments passed to or from other methods
Details
This implementation is based on lm.gls and errorsarlm. In particular, the function does not (yet) prevent asymmetric spatial weights being used with "CAR" family models. It appears that both numerical issues (convergence in particular) and uncertainties about the exact spatial weights matrix used make it difficult to reproduce Cressie and Chan's 1989 results, also given in Cressie 1993.
Note that the fitted() function for the output object assumes that the response variable may be reconstructed as the sum of the trend, the signal, and the noise (residuals). Since the values of the response variable are known, their spatial lags are used to calculate signal components (Cressie 1993, p. 564). This differs from other software, including GeoDa, which does not use knowledge of the response variable in making predictions for the fitting data.
Control arguments
- tol.opt:
the desired accuracy of the optimization - passed to
optimize()(default=.Machine$double.eps^(2/3))- fdHess:
default NULL, then set to (method != "eigen") internally; use
fdHessto compute an approximate Hessian using finite differences when using sparse matrix methods; used to make a coefficient covariance matrix when the number of observations is large; may be turned off to save resources if need be- optimHess:
default FALSE, use
fdHessfrom nlme, if TRUE, useoptimto calculate Hessian at optimum- optimHessMethod:
default “optimHess”, may be “nlm” or one of the
optimmethods- Imult:
default 2; used for preparing the Cholesky decompositions for updating in the Jacobian function
- super:
if NULL (default), set to FALSE to use a simplicial decomposition for the sparse Cholesky decomposition and method “Matrix_J”, set to
as.logical(NA)for method “Matrix”, if TRUE, use a supernodal decomposition- cheb_q:
default 5; highest power of the approximating polynomial for the Chebyshev approximation
- MC_p:
default 16; number of random variates
- MC_m:
default 30; number of products of random variates matrix and spatial weights matrix
- type
default “MC”, used with method “moments”; alternatives “mult” and “moments”, for use if
trsis missing,trW- correct
default TRUE, used with method “moments” to compute the Smirnov/Anselin correction term
- trunc
default TRUE, used with method “moments” to truncate the Smirnov/Anselin correction term
- SE_method
default “LU”, may be “MC”
- nrho
default 200, as in SE toolbox; the size of the first stage lndet grid; it may be reduced to for example 40
- interpn
default 2000, as in SE toolbox; the size of the second stage lndet grid
- small_asy
default TRUE; if the method is not “eigen”, use asymmetric covariances rather than numerical Hessian ones if n <= small
- small
default 1500; threshold number of observations for asymmetric covariances when the method is not “eigen”
- SElndet
default NULL, may be used to pass a pre-computed SE toolbox style matrix of coefficients and their lndet values to the "SE_classic" and "SE_whichMin" methods
- LU_order
default FALSE; used in “LU_prepermutate”, note warnings given for
lumethod- pre_eig
default NULL; may be used to pass a pre-computed vector of eigenvalues
Value
A list object of class Spautolm:
- fit
a list, with items:
- coefficients
ML coefficient estimates
- SSE
ML sum of squared errors
- s2
ML residual variance
- imat
ML coefficient covariance matrix (before multiplying by s2)
- signal_trend
non-spatial component of fitted.values
- signal_stochastic
spatial component of fitted.values
- fitted.values
sum of non-spatial and spatial components of fitted.values
- residuals
difference between observed and fitted values
- lambda
ML autoregressive coefficient
- LL
log likelihood for fitted model
- LL0
log likelihood for model with lambda=0
- call
the call used to create this object
- parameters
number of parameters estimated
- aliased
if not NULL, details of aliased variables
- method
Jacobian method chosen
- family
family chosen
- zero.policy
zero.policy used
- weights
case weights used
- interval
the line search interval used
- timings
processing timings
- na.action
(possibly) named vector of excluded or omitted observations if non-default na.action argument used
- llprof
if not NULL, a list with components lambda and ll of equal length
- lambda.se
Numerical Hessian-based standard error of lambda
- fdHess
Numerical Hessian-based variance-covariance matrix
- X
covariates used in model fitting
- Y
response used in model fitting
- weights
weights used in model fitting
References
Cliff, A. D., Ord, J. K. 1981 Spatial processes, Pion; Ord, J. K. 1975 Estimation methods for models of spatial interaction, Journal of the American Statistical Association, 70, 120-126; Waller, L. A., Gotway, C. A. 2004 Applied spatial statistics for public health, Wiley, Hoboken, NJ, 325-380; Cressie, N. A. C. 1993 Statistics for spatial data, Wiley, New York, 548-568; Ripley, B. D. 1981 Spatial statistics, Wiley, New York, 88-95; LeSage J and RK Pace (2009) Introduction to Spatial Econometrics. CRC Press, Boca Raton.
Author
Roger Bivand Roger.Bivand@nhh.no
Note
The standard errors given in Waller and Gotway (2004) are adjusted for the numbers of parameters estimated, and may be reproduced by using the additional argument adj.se=TRUE in the summary method. In addition, the function returns fitted values and residuals as given by Cressie (1993) p. 564.
Examples
require("sf", quietly=TRUE)
nydata <- st_read(system.file("shapes/NY8_bna_utm18.gpkg", package="spData")[1], quiet=TRUE)
# \dontrun{
lm0 <- lm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata)
summary(lm0)
#>
#> Call:
#> lm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.7417 -0.3957 -0.0326 0.3353 4.1398
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.51728 0.15856 -3.262 0.00124 **
#> PEXPOSURE 0.04884 0.03506 1.393 0.16480
#> PCTAGE65P 3.95089 0.60550 6.525 3.22e-10 ***
#> PCTOWNHOME -0.56004 0.17031 -3.288 0.00114 **
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#>
#> Residual standard error: 0.6571 on 277 degrees of freedom
#> Multiple R-squared: 0.1932, Adjusted R-squared: 0.1844
#> F-statistic: 22.1 on 3 and 277 DF, p-value: 7.306e-13
#>
lm0w <- lm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata, weights=POP8)
summary(lm0w)
#>
#> Call:
#> lm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> weights = POP8)
#>
#> Weighted Residuals:
#> Min 1Q Median 3Q Max
#> -129.067 -14.714 5.817 25.624 70.723
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) -0.77837 0.14116 -5.514 8.03e-08 ***
#> PEXPOSURE 0.07626 0.02731 2.792 0.00560 **
#> PCTAGE65P 3.85656 0.57126 6.751 8.60e-11 ***
#> PCTOWNHOME -0.39869 0.15305 -2.605 0.00968 **
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#>
#> Residual standard error: 33.5 on 277 degrees of freedom
#> Multiple R-squared: 0.1977, Adjusted R-squared: 0.189
#> F-statistic: 22.75 on 3 and 277 DF, p-value: 3.382e-13
#>
# }
suppressMessages(nyadjmat <- as.matrix(foreign::read.dbf(system.file(
"misc/nyadjwts.dbf", package="spData")[1])[-1]))
suppressMessages(ID <- as.character(names(foreign::read.dbf(system.file(
"misc/nyadjwts.dbf", package="spData")[1]))[-1]))
identical(substring(ID, 2, 10), substring(as.character(nydata$AREAKEY), 2, 10))
#> [1] TRUE
#require("spdep", quietly=TRUE)
listw_NY <- spdep::mat2listw(nyadjmat, as.character(nydata$AREAKEY), style="B")
eigs <- eigenw(listw_NY)
# \dontrun{
esar0 <- errorsarlm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata,
listw=listw_NY)
summary(esar0)
#>
#> Call:errorsarlm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME,
#> data = nydata, listw = listw_NY)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.56754 -0.38239 -0.02643 0.33109 4.01219
#>
#> Type: error
#> Coefficients: (asymptotic standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.618193 0.176784 -3.4969 0.0004707
#> PEXPOSURE 0.071014 0.042051 1.6888 0.0912635
#> PCTAGE65P 3.754200 0.624722 6.0094 1.862e-09
#> PCTOWNHOME -0.419890 0.191329 -2.1946 0.0281930
#>
#> Lambda: 0.040487, LR test value: 5.2438, p-value: 0.022026
#> Asymptotic standard error: 0.016214
#> z-value: 2.4971, p-value: 0.01252
#> Wald statistic: 6.2356, p-value: 0.01252
#>
#> Log likelihood: -276.1069 for error model
#> ML residual variance (sigma squared): 0.41388, (sigma: 0.64333)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: 564.21, (AIC for lm: 567.46)
#>
system.time(esar1f <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME,
data=nydata, listw=listw_NY, family="SAR", method="eigen",
control=list(pre_eig=eigs)))
#> user system elapsed
#> 0.220 0.000 0.222
res <- summary(esar1f)
print(res)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, family = "SAR", method = "eigen", control = list(pre_eig = eigs))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.56754 -0.38239 -0.02643 0.33109 4.01219
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.618193 0.176784 -3.4969 0.0004707
#> PEXPOSURE 0.071014 0.042051 1.6888 0.0912635
#> PCTAGE65P 3.754200 0.624722 6.0094 1.862e-09
#> PCTOWNHOME -0.419890 0.191329 -2.1946 0.0281930
#>
#> Lambda: 0.040487 LR test value: 5.2438 p-value: 0.022026
#> Numerical Hessian standard error of lambda: 0.017197
#>
#> Log likelihood: -276.1069
#> ML residual variance (sigma squared): 0.41388, (sigma: 0.64333)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: 564.21
#>
coef(res)
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.61819272 0.17678351 -3.496891 4.707136e-04
#> PEXPOSURE 0.07101384 0.04205063 1.688770 9.126350e-02
#> PCTAGE65P 3.75419996 0.62472153 6.009397 1.862142e-09
#> PCTOWNHOME -0.41988960 0.19132936 -2.194591 2.819298e-02
sqrt(diag(res$resvar))
#> (Intercept) PEXPOSURE PCTAGE65P PCTOWNHOME
#> 0.17678351 0.04205063 0.62472153 0.19132936
sqrt(diag(esar1f$fit$imat)*esar1f$fit$s2)
#> (Intercept) PEXPOSURE PCTAGE65P PCTOWNHOME
#> 0.17678351 0.04205063 0.62472153 0.19132936
sqrt(diag(esar1f$fdHess))
#> [1] 0.01719720 0.18518821 0.04385029 0.62998618 0.20355594
system.time(esar1M <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME,
data=nydata, listw=listw_NY, family="SAR", method="Matrix"))
#> user system elapsed
#> 0.243 0.000 0.246
summary(esar1M)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, family = "SAR", method = "Matrix")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -3.406132 -0.561646 -0.092662 0.474796 5.384405
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.414826 0.102166 -4.0603 4.901e-05
#> PEXPOSURE 0.015081 0.017772 0.8486 0.3961
#> PCTAGE65P 5.159749 0.476498 10.8285 < 2.2e-16
#> PCTOWNHOME -0.892387 0.099241 -8.9921 < 2.2e-16
#>
#> Lambda: -0.38889 LR test value: 254.73 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: 0.044856
#>
#> Log likelihood: -151.3662
#> ML residual variance (sigma squared): 0.95111, (sigma: 0.97525)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: 314.73
#>
system.time(esar1M <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME,
data=nydata, listw=listw_NY, family="SAR", method="Matrix",
control=list(super=TRUE)))
#> user system elapsed
#> 0.225 0.000 0.226
summary(esar1M)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, family = "SAR", method = "Matrix", control = list(super = TRUE))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -3.178535 -0.521860 -0.074421 0.414212 5.184465
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.411041 0.104426 -3.9362 8.279e-05
#> PEXPOSURE 0.015768 0.018366 0.8585 0.3906
#> PCTAGE65P 5.070130 0.483332 10.4900 < 2.2e-16
#> PCTOWNHOME -0.880883 0.102093 -8.6282 < 2.2e-16
#>
#> Lambda: -0.33146 LR test value: 247.44 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: 0.030656
#>
#> Log likelihood: -155.0108
#> ML residual variance (sigma squared): 0.82436, (sigma: 0.90795)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: 322.02
#>
esar1wf <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata,
listw=listw_NY, weights=POP8, family="SAR", method="eigen",
control=list(pre_eig=eigs))
summary(esar1wf)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, weights = POP8, family = "SAR", method = "eigen",
#> control = list(pre_eig = eigs))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.48488 -0.26823 0.09489 0.46552 4.28343
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.797063 0.144054 -5.5331 3.146e-08
#> PEXPOSURE 0.080545 0.028334 2.8428 0.004473
#> PCTAGE65P 3.816731 0.576037 6.6258 3.453e-11
#> PCTOWNHOME -0.380778 0.156507 -2.4330 0.014975
#>
#> Lambda: 0.0095636 LR test value: 0.32665 p-value: 0.56764
#> Numerical Hessian standard error of lambda: 0.016356
#>
#> Log likelihood: -251.6017
#> ML residual variance (sigma squared): 1104.1, (sigma: 33.229)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: NA (not available for weighted model)
#>
system.time(esar1wM <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME,
data=nydata, listw=listw_NY, weights=POP8, family="SAR", method="Matrix"))
#> user system elapsed
#> 0.241 0.000 0.243
summary(esar1wM)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, weights = POP8, family = "SAR", method = "Matrix")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.561100 -0.374524 0.057405 0.591094 5.700142
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.578546 0.090006 -6.4279 1.294e-10
#> PEXPOSURE 0.035402 0.013959 2.5361 0.01121
#> PCTAGE65P 4.651137 0.421285 11.0404 < 2.2e-16
#> PCTOWNHOME -0.666898 0.091443 -7.2931 3.029e-13
#>
#> Lambda: -0.34423 LR test value: 264.24 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: 0.036784
#>
#> Log likelihood: -119.6468
#> ML residual variance (sigma squared): 2129.1, (sigma: 46.142)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: NA (not available for weighted model)
#>
esar1wlu <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata,
listw=listw_NY, weights=POP8, family="SAR", method="LU")
summary(esar1wlu)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, weights = POP8, family = "SAR", method = "LU")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.48488 -0.26823 0.09489 0.46552 4.28343
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.797063 0.144054 -5.5331 3.146e-08
#> PEXPOSURE 0.080545 0.028334 2.8428 0.004473
#> PCTAGE65P 3.816731 0.576037 6.6258 3.453e-11
#> PCTOWNHOME -0.380778 0.156507 -2.4330 0.014975
#>
#> Lambda: 0.0095636 LR test value: 0.32665 p-value: 0.56764
#> Numerical Hessian standard error of lambda: 0.016623
#>
#> Log likelihood: -251.6017
#> ML residual variance (sigma squared): 1104.1, (sigma: 33.229)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: NA (not available for weighted model)
#>
esar1wch <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata,
listw=listw_NY, weights=POP8, family="SAR", method="Chebyshev")
summary(esar1wch)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, weights = POP8, family = "SAR", method = "Chebyshev")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -4.39831 -0.86303 0.12117 1.09320 8.78570
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.538555 0.087573 -6.1498 7.76e-10
#> PEXPOSURE 0.029782 0.013074 2.2780 0.02273
#> PCTAGE65P 4.896346 0.419359 11.6758 < 2.2e-16
#> PCTOWNHOME -0.737829 0.087569 -8.4257 < 2.2e-16
#>
#> Lambda: -1 LR test value: 236970 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: NaN
#>
#> Log likelihood: 118232.5
#> ML residual variance (sigma squared): 9336.4, (sigma: 96.625)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: NA (not available for weighted model)
#>
# }
ecar1f <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata,
listw=listw_NY, family="CAR", method="eigen",
control=list(pre_eig=eigs))
summary(ecar1f)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, family = "CAR", method = "eigen", control = list(pre_eig = eigs))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.539732 -0.384311 -0.030646 0.335126 3.808848
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.648362 0.181129 -3.5796 0.0003442
#> PEXPOSURE 0.077899 0.043692 1.7829 0.0745986
#> PCTAGE65P 3.703830 0.627185 5.9055 3.516e-09
#> PCTOWNHOME -0.382789 0.195564 -1.9574 0.0503053
#>
#> Lambda: 0.084123 LR test value: 5.8009 p-value: 0.016018
#> Numerical Hessian standard error of lambda: 0.030853
#>
#> Log likelihood: -275.8283
#> ML residual variance (sigma squared): 0.40758, (sigma: 0.63842)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: 563.66
#>
# \dontrun{
system.time(ecar1M <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME,
data=nydata, listw=listw_NY, family="CAR", method="Matrix"))
#> user system elapsed
#> 0.274 0.000 0.275
summary(ecar1M)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, family = "CAR", method = "Matrix")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -3.449951 -0.633777 -0.072436 0.550248 6.039594
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.402788 0.112612 -3.5768 0.0003479
#> PEXPOSURE 0.020131 0.020783 0.9686 0.3327222
#> PCTAGE65P 4.632644 0.500526 9.2555 < 2.2e-16
#> PCTOWNHOME -0.812981 0.112867 -7.2030 5.891e-13
#>
#> Lambda: -0.5 LR test value: 228.48 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: NaN
#>
#> Log likelihood: -164.4897
#> ML residual variance (sigma squared): 0.50386, (sigma: 0.70983)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: 340.98
#>
# }
ecar1wf <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data=nydata,
listw=listw_NY, weights=POP8, family="CAR", method="eigen",
control=list(pre_eig=eigs))
summary(ecar1wf)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, weights = POP8, family = "CAR", method = "eigen",
#> control = list(pre_eig = eigs))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.491042 -0.270906 0.081435 0.451556 4.198134
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.790154 0.144862 -5.4545 4.910e-08
#> PEXPOSURE 0.081922 0.028593 2.8651 0.004169
#> PCTAGE65P 3.825858 0.577720 6.6223 3.536e-11
#> PCTOWNHOME -0.386820 0.157436 -2.4570 0.014010
#>
#> Lambda: 0.022419 LR test value: 0.38785 p-value: 0.53343
#> Numerical Hessian standard error of lambda: 0.038916
#>
#> Log likelihood: -251.5711
#> ML residual variance (sigma squared): 1102.9, (sigma: 33.21)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: NA (not available for weighted model)
#>
# \dontrun{
system.time(ecar1wM <- spautolm(Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME,
data=nydata, listw=listw_NY, weights=POP8, family="CAR", method="Matrix"))
#> user system elapsed
#> 0.274 0.000 0.276
summary(ecar1wM)
#>
#> Call:
#> spautolm(formula = Z ~ PEXPOSURE + PCTAGE65P + PCTOWNHOME, data = nydata,
#> listw = listw_NY, weights = POP8, family = "CAR", method = "Matrix")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.98144 -0.15716 0.37342 1.08857 7.10495
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.714952 0.093905 -7.6136 2.665e-14
#> PEXPOSURE 0.041467 0.015279 2.7139 0.00665
#> PCTAGE65P 4.149207 0.425446 9.7526 < 2.2e-16
#> PCTOWNHOME -0.478396 0.097030 -4.9304 8.205e-07
#>
#> Lambda: -0.5 LR test value: 262.65 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: NaN
#>
#> Log likelihood: -120.4395
#> ML residual variance (sigma squared): 1159.2, (sigma: 34.046)
#> Number of observations: 281
#> Number of parameters estimated: 6
#> AIC: NA (not available for weighted model)
#>
# }
# \dontrun{
require("sf", quietly=TRUE)
nc.sids <- st_read(system.file("shapes/sids.gpkg", package="spData")[1], quiet=TRUE)
ft.SID74 <- sqrt(1000)*(sqrt(nc.sids$SID74/nc.sids$BIR74) +
sqrt((nc.sids$SID74+1)/nc.sids$BIR74))
lm_nc <- lm(ft.SID74 ~ 1)
sids.nhbr30 <- spdep::dnearneigh(cbind(nc.sids$east, nc.sids$north), 0, 30,
row.names=row.names(nc.sids))
#> Warning: neighbour object has 3 sub-graphs
sids.nhbr30.dist <- spdep::nbdists(sids.nhbr30, cbind(nc.sids$east, nc.sids$north))
sids.nhbr <- spdep::listw2sn(spdep::nb2listw(sids.nhbr30,
glist=sids.nhbr30.dist, style="B", zero.policy=TRUE))
dij <- sids.nhbr[,3]
n <- nc.sids$BIR74
el1 <- min(dij)/dij
el2 <- sqrt(n[sids.nhbr$to]/n[sids.nhbr$from])
sids.nhbr$weights <- el1*el2
sids.nhbr.listw <- spdep::sn2listw(sids.nhbr, style="B", zero.policy=TRUE)
#> Warning: no-neighbour observations found, set zero.policy to TRUE;
#> this warning will soon become an error
#> Warning: neighbour object has 3 sub-graphs
both <- factor(paste(nc.sids$L_id, nc.sids$M_id, sep=":"))
ft.NWBIR74 <- sqrt(1000)*(sqrt(nc.sids$NWBIR74/nc.sids$BIR74) +
sqrt((nc.sids$NWBIR74+1)/nc.sids$BIR74))
mdata <- data.frame(both, ft.NWBIR74, ft.SID74, BIR74=nc.sids$BIR74)
outl <- which.max(rstandard(lm_nc))
as.character(nc.sids$NAME[outl])
#> [1] "Anson"
mdata.4 <- mdata[-outl,]
W <- spdep::listw2mat(sids.nhbr.listw)
W.4 <- W[-outl, -outl]
sids.nhbr.listw.4 <- spdep::mat2listw(W.4, style="B", zero.policy=TRUE)
#> Warning: neighbour object has 3 sub-graphs
esarI <- errorsarlm(ft.SID74 ~ 1, data=mdata, listw=sids.nhbr.listw,
zero.policy=TRUE)
summary(esarI)
#>
#> Call:errorsarlm(formula = ft.SID74 ~ 1, data = mdata, listw = sids.nhbr.listw,
#> zero.policy = TRUE)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.887117 -0.636573 -0.043429 0.448767 3.406724
#>
#> Type: error
#> Regions with no neighbours included:
#> 56 87
#> Coefficients: (asymptotic standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 2.97463 0.13011 22.862 < 2.2e-16
#>
#> Lambda: 0.66864, LR test value: 10.214, p-value: 0.0013939
#> Asymptotic standard error: 0.11473
#> z-value: 5.8279, p-value: 5.6145e-09
#> Wald statistic: 33.964, p-value: 5.6145e-09
#>
#> Log likelihood: -133.8616 for error model
#> ML residual variance (sigma squared): 0.81932, (sigma: 0.90516)
#> Number of observations: 100
#> Number of parameters estimated: 3
#> AIC: 273.72, (AIC for lm: 281.94)
#>
esarIa <- spautolm(ft.SID74 ~ 1, data=mdata, listw=sids.nhbr.listw,
family="SAR")
summary(esarIa)
#>
#> Call: spautolm(formula = ft.SID74 ~ 1, data = mdata, listw = sids.nhbr.listw,
#> family = "SAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.887117 -0.636573 -0.043429 0.448767 3.406724
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 2.97463 0.13011 22.862 < 2.2e-16
#>
#> Lambda: 0.66864 LR test value: 10.214 p-value: 0.0013939
#> Numerical Hessian standard error of lambda: 0.16505
#>
#> Log likelihood: -133.8616
#> ML residual variance (sigma squared): 0.81932, (sigma: 0.90516)
#> Number of observations: 100
#> Number of parameters estimated: 3
#> AIC: 273.72
#>
esarIV <- errorsarlm(ft.SID74 ~ ft.NWBIR74, data=mdata, listw=sids.nhbr.listw,
zero.policy=TRUE)
summary(esarIV)
#>
#> Call:
#> errorsarlm(formula = ft.SID74 ~ ft.NWBIR74, data = mdata, listw = sids.nhbr.listw,
#> zero.policy = TRUE)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.123648 -0.573163 0.017859 0.468022 2.693604
#>
#> Type: error
#> Regions with no neighbours included:
#> 56 87
#> Coefficients: (asymptotic standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 1.549443 0.219230 7.0677 1.576e-12
#> ft.NWBIR74 0.041974 0.006171 6.8018 1.033e-11
#>
#> Lambda: 0.18465, LR test value: 0.50496, p-value: 0.47733
#> Asymptotic standard error: 0.20648
#> z-value: 0.89424, p-value: 0.37119
#> Wald statistic: 0.79967, p-value: 0.37119
#>
#> Log likelihood: -117.7464 for error model
#> ML residual variance (sigma squared): 0.61546, (sigma: 0.78451)
#> Number of observations: 100
#> Number of parameters estimated: 4
#> AIC: 243.49, (AIC for lm: 242)
#>
esarIVa <- spautolm(ft.SID74 ~ ft.NWBIR74, data=mdata, listw=sids.nhbr.listw,
family="SAR")
summary(esarIVa)
#>
#> Call:
#> spautolm(formula = ft.SID74 ~ ft.NWBIR74, data = mdata, listw = sids.nhbr.listw,
#> family = "SAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.123648 -0.573163 0.017859 0.468022 2.693604
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 1.549443 0.219230 7.0677 1.576e-12
#> ft.NWBIR74 0.041974 0.006171 6.8018 1.033e-11
#>
#> Lambda: 0.18465 LR test value: 0.50496 p-value: 0.47733
#> Numerical Hessian standard error of lambda: 0.25601
#>
#> Log likelihood: -117.7464
#> ML residual variance (sigma squared): 0.61546, (sigma: 0.78451)
#> Number of observations: 100
#> Number of parameters estimated: 4
#> AIC: 243.49
#>
esarIaw <- spautolm(ft.SID74 ~ 1, data=mdata, listw=sids.nhbr.listw,
weights=BIR74, family="SAR")
summary(esarIaw)
#>
#> Call: spautolm(formula = ft.SID74 ~ 1, data = mdata, listw = sids.nhbr.listw,
#> weights = BIR74, family = "SAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.867485 -0.568644 0.019717 0.502197 3.498013
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 2.852052 0.090271 31.594 < 2.2e-16
#>
#> Lambda: 0.7338 LR test value: 12.917 p-value: 0.00032554
#> Numerical Hessian standard error of lambda: 0.13886
#>
#> Log likelihood: -130.0975
#> ML residual variance (sigma squared): 1539.4, (sigma: 39.236)
#> Number of observations: 100
#> Number of parameters estimated: 3
#> AIC: NA (not available for weighted model)
#>
esarIIaw <- spautolm(ft.SID74 ~ both - 1, data=mdata, listw=sids.nhbr.listw,
weights=BIR74, family="SAR")
summary(esarIIaw)
#>
#> Call:
#> spautolm(formula = ft.SID74 ~ both - 1, data = mdata, listw = sids.nhbr.listw,
#> weights = BIR74, family = "SAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.590809 -0.432976 0.016736 0.357284 3.536718
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> both1:2 2.05545 0.22184 9.2654 < 2.2e-16
#> both1:3 2.87260 0.16181 17.7531 < 2.2e-16
#> both1:4 4.16365 0.34330 12.1283 < 2.2e-16
#> both2:1 2.47255 0.29757 8.3090 < 2.2e-16
#> both2:2 2.15307 0.21172 10.1692 < 2.2e-16
#> both2:3 2.64235 0.17296 15.2770 < 2.2e-16
#> both2:4 3.26604 0.28287 11.5459 < 2.2e-16
#> both3:1 3.11277 0.34166 9.1107 < 2.2e-16
#> both3:2 2.76541 0.15667 17.6508 < 2.2e-16
#> both3:3 2.86582 0.18593 15.4134 < 2.2e-16
#> both3:4 3.18142 0.21617 14.7169 < 2.2e-16
#> both4:3 3.69333 0.23348 15.8188 < 2.2e-16
#>
#> Lambda: 0.32136 LR test value: 1.4004 p-value: 0.23666
#> Numerical Hessian standard error of lambda: 0.25501
#>
#> Log likelihood: -109.8922
#> ML residual variance (sigma squared): 1071.6, (sigma: 32.735)
#> Number of observations: 100
#> Number of parameters estimated: 14
#> AIC: NA (not available for weighted model)
#>
esarIVaw <- spautolm(ft.SID74 ~ ft.NWBIR74, data=mdata,
listw=sids.nhbr.listw, weights=BIR74, family="SAR")
summary(esarIVaw)
#>
#> Call:
#> spautolm(formula = ft.SID74 ~ ft.NWBIR74, data = mdata, listw = sids.nhbr.listw,
#> weights = BIR74, family = "SAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.00956 -0.45229 0.12547 0.55952 2.92223
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 1.5769279 0.2501334 6.3043 2.894e-10
#> ft.NWBIR74 0.0368573 0.0069413 5.3099 1.097e-07
#>
#> Lambda: 0.3839 LR test value: 1.9983 p-value: 0.15747
#> Numerical Hessian standard error of lambda: 0.25778
#>
#> Log likelihood: -119.5648
#> ML residual variance (sigma squared): 1295.8, (sigma: 35.997)
#> Number of observations: 100
#> Number of parameters estimated: 4
#> AIC: NA (not available for weighted model)
#>
ecarIaw <- spautolm(ft.SID74 ~ 1, data=mdata.4, listw=sids.nhbr.listw.4,
weights=BIR74, family="CAR")
#> Warning: Non-symmetric spatial weights in CAR model
summary(ecarIaw)
#>
#> Call:
#> spautolm(formula = ft.SID74 ~ 1, data = mdata.4, listw = sids.nhbr.listw.4,
#> weights = BIR74, family = "CAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.009350 -0.638915 -0.060761 0.428526 2.019409
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 2.942864 0.095304 30.879 < 2.2e-16
#>
#> Lambda: 0.86832 LR test value: 23.003 p-value: 1.6172e-06
#> Numerical Hessian standard error of lambda: 0.048102
#>
#> Log likelihood: -118.7564
#> ML residual variance (sigma squared): 1264, (sigma: 35.553)
#> Number of observations: 99
#> Number of parameters estimated: 3
#> AIC: NA (not available for weighted model)
#>
ecarIIaw <- spautolm(ft.SID74 ~ both - 1, data=mdata.4,
listw=sids.nhbr.listw.4, weights=BIR74, family="CAR")
#> Warning: Non-symmetric spatial weights in CAR model
#> Warning: NaNs produced
summary(ecarIIaw)
#>
#> Call:
#> spautolm(formula = ft.SID74 ~ both - 1, data = mdata.4, listw = sids.nhbr.listw.4,
#> weights = BIR74, family = "CAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.564067 -0.461531 -0.020982 0.384458 2.054255
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> both1:2 2.06282 0.20065 10.2806 < 2.2e-16
#> both1:3 2.91982 0.14171 20.6048 < 2.2e-16
#> both1:4 4.12159 0.30076 13.7037 < 2.2e-16
#> both2:1 2.58281 0.27014 9.5611 < 2.2e-16
#> both2:2 2.17549 0.18265 11.9104 < 2.2e-16
#> both2:3 2.67030 0.15355 17.3910 < 2.2e-16
#> both2:4 3.10806 0.24748 12.5588 < 2.2e-16
#> both3:1 2.93237 0.30007 9.7724 < 2.2e-16
#> both3:2 2.65317 0.14139 18.7646 < 2.2e-16
#> both3:3 2.91685 0.17134 17.0234 < 2.2e-16
#> both3:4 3.20447 0.20402 15.7063 < 2.2e-16
#> both4:3 3.80672 0.20831 18.2742 < 2.2e-16
#>
#> Lambda: 0.22163 LR test value: 1.3827 p-value: 0.23964
#> Numerical Hessian standard error of lambda: NaN
#>
#> Log likelihood: -99.2181
#> ML residual variance (sigma squared): 890.66, (sigma: 29.844)
#> Number of observations: 99
#> Number of parameters estimated: 14
#> AIC: NA (not available for weighted model)
#>
ecarIVaw <- spautolm(ft.SID74 ~ ft.NWBIR74, data=mdata.4,
listw=sids.nhbr.listw.4, weights=BIR74, family="CAR")
#> Warning: Non-symmetric spatial weights in CAR model
summary(ecarIVaw)
#>
#> Call:
#> spautolm(formula = ft.SID74 ~ ft.NWBIR74, data = mdata.4, listw = sids.nhbr.listw.4,
#> weights = BIR74, family = "CAR")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.99259 -0.44794 0.15464 0.60748 1.95751
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 1.434705 0.225521 6.3618 1.995e-10
#> ft.NWBIR74 0.040903 0.006299 6.4936 8.382e-11
#>
#> Lambda: 0.22724 LR test value: 1.1936 p-value: 0.2746
#> Numerical Hessian standard error of lambda: 0.55239
#>
#> Log likelihood: -114.0196
#> ML residual variance (sigma squared): 1201, (sigma: 34.655)
#> Number of observations: 99
#> Number of parameters estimated: 4
#> AIC: NA (not available for weighted model)
#>
nc.sids$fitIV <- append(fitted.values(ecarIVaw), NA, outl-1)
plot(nc.sids[,"fitIV"], nbreaks=12) # Cressie 1993, p. 565
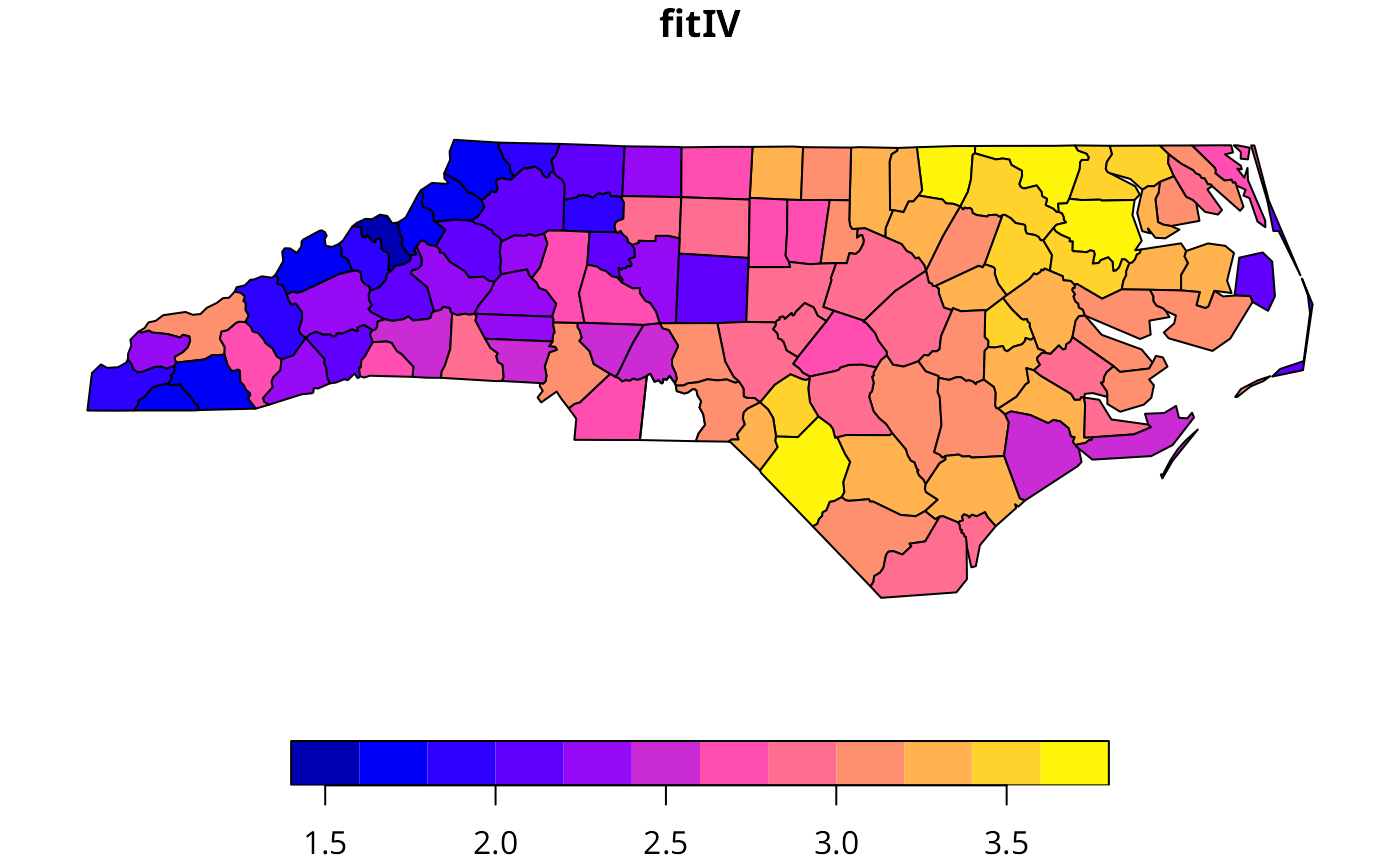 # }
# \dontrun{
data(oldcol, package="spdep")
COL.errW.eig <- errorsarlm(CRIME ~ INC + HOVAL, data=COL.OLD,
spdep::nb2listw(COL.nb, style="W"))
summary(COL.errW.eig)
#>
#> Call:
#> errorsarlm(formula = CRIME ~ INC + HOVAL, data = COL.OLD, listw = spdep::nb2listw(COL.nb,
#> style = "W"))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -34.81174 -6.44031 -0.72142 7.61476 23.33626
#>
#> Type: error
#> Coefficients: (asymptotic standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 59.893219 5.366163 11.1613 < 2.2e-16
#> INC -0.941312 0.330569 -2.8476 0.0044057
#> HOVAL -0.302250 0.090476 -3.3407 0.0008358
#>
#> Lambda: 0.56179, LR test value: 7.9935, p-value: 0.0046945
#> Asymptotic standard error: 0.13387
#> z-value: 4.1966, p-value: 2.7098e-05
#> Wald statistic: 17.611, p-value: 2.7098e-05
#>
#> Log likelihood: -183.3805 for error model
#> ML residual variance (sigma squared): 95.575, (sigma: 9.7762)
#> Number of observations: 49
#> Number of parameters estimated: 5
#> AIC: 376.76, (AIC for lm: 382.75)
#>
COL.errW.sar <- spautolm(CRIME ~ INC + HOVAL, data=COL.OLD,
spdep::nb2listw(COL.nb, style="W"))
summary(COL.errW.sar)
#>
#> Call:
#> spautolm(formula = CRIME ~ INC + HOVAL, data = COL.OLD, listw = spdep::nb2listw(COL.nb,
#> style = "W"))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -34.81174 -6.44031 -0.72142 7.61476 23.33626
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 59.893219 5.366163 11.1613 < 2.2e-16
#> INC -0.941312 0.330569 -2.8476 0.0044057
#> HOVAL -0.302250 0.090476 -3.3407 0.0008358
#>
#> Lambda: 0.56179 LR test value: 7.9935 p-value: 0.0046945
#> Numerical Hessian standard error of lambda: 0.15242
#>
#> Log likelihood: -183.3805
#> ML residual variance (sigma squared): 95.575, (sigma: 9.7762)
#> Number of observations: 49
#> Number of parameters estimated: 5
#> AIC: 376.76
#>
data(boston, package="spData")
gp1 <- spautolm(log(CMEDV) ~ CRIM + ZN + INDUS + CHAS + I(NOX^2)
+ I(RM^2) + AGE + log(DIS) + log(RAD) + TAX + PTRATIO + B + log(LSTAT),
data=boston.c, spdep::nb2listw(boston.soi), family="SMA")
summary(gp1)
#>
#> Call: spautolm(formula = log(CMEDV) ~ CRIM + ZN + INDUS + CHAS + I(NOX^2) +
#> I(RM^2) + AGE + log(DIS) + log(RAD) + TAX + PTRATIO + B +
#> log(LSTAT), data = boston.c, listw = spdep::nb2listw(boston.soi),
#> family = "SMA")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -0.5847694 -0.0713881 0.0012284 0.0827517 0.6071219
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 4.28501607 0.15367176 27.8842 < 2.2e-16
#> CRIM -0.00718807 0.00106298 -6.7622 1.359e-11
#> ZN 0.00023008 0.00051897 0.4433 0.6575185
#> INDUS 0.00047300 0.00263339 0.1796 0.8574551
#> CHAS1 0.01020698 0.02872047 0.3554 0.7222970
#> I(NOX^2) -0.44885530 0.13675913 -3.2821 0.0010304
#> I(RM^2) 0.00638094 0.00110330 5.7835 7.316e-09
#> AGE -0.00043973 0.00051336 -0.8566 0.3916862
#> log(DIS) -0.15650578 0.03856337 -4.0584 4.941e-05
#> log(RAD) 0.07583760 0.02016468 3.7609 0.0001693
#> TAX -0.00049364 0.00012162 -4.0588 4.933e-05
#> PTRATIO -0.02494959 0.00538791 -4.6307 3.645e-06
#> B 0.00048517 0.00010944 4.4334 9.277e-06
#> log(LSTAT) -0.32961379 0.02353891 -14.0029 < 2.2e-16
#>
#> Lambda: 0.61991 LR test value: 144.28 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: 0.04431
#>
#> Log likelihood: 229.1208
#> ML residual variance (sigma squared): 0.02596, (sigma: 0.16112)
#> Number of observations: 506
#> Number of parameters estimated: 16
#> AIC: -426.24
#>
# }
# }
# \dontrun{
data(oldcol, package="spdep")
COL.errW.eig <- errorsarlm(CRIME ~ INC + HOVAL, data=COL.OLD,
spdep::nb2listw(COL.nb, style="W"))
summary(COL.errW.eig)
#>
#> Call:
#> errorsarlm(formula = CRIME ~ INC + HOVAL, data = COL.OLD, listw = spdep::nb2listw(COL.nb,
#> style = "W"))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -34.81174 -6.44031 -0.72142 7.61476 23.33626
#>
#> Type: error
#> Coefficients: (asymptotic standard errors)
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 59.893219 5.366163 11.1613 < 2.2e-16
#> INC -0.941312 0.330569 -2.8476 0.0044057
#> HOVAL -0.302250 0.090476 -3.3407 0.0008358
#>
#> Lambda: 0.56179, LR test value: 7.9935, p-value: 0.0046945
#> Asymptotic standard error: 0.13387
#> z-value: 4.1966, p-value: 2.7098e-05
#> Wald statistic: 17.611, p-value: 2.7098e-05
#>
#> Log likelihood: -183.3805 for error model
#> ML residual variance (sigma squared): 95.575, (sigma: 9.7762)
#> Number of observations: 49
#> Number of parameters estimated: 5
#> AIC: 376.76, (AIC for lm: 382.75)
#>
COL.errW.sar <- spautolm(CRIME ~ INC + HOVAL, data=COL.OLD,
spdep::nb2listw(COL.nb, style="W"))
summary(COL.errW.sar)
#>
#> Call:
#> spautolm(formula = CRIME ~ INC + HOVAL, data = COL.OLD, listw = spdep::nb2listw(COL.nb,
#> style = "W"))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -34.81174 -6.44031 -0.72142 7.61476 23.33626
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 59.893219 5.366163 11.1613 < 2.2e-16
#> INC -0.941312 0.330569 -2.8476 0.0044057
#> HOVAL -0.302250 0.090476 -3.3407 0.0008358
#>
#> Lambda: 0.56179 LR test value: 7.9935 p-value: 0.0046945
#> Numerical Hessian standard error of lambda: 0.15242
#>
#> Log likelihood: -183.3805
#> ML residual variance (sigma squared): 95.575, (sigma: 9.7762)
#> Number of observations: 49
#> Number of parameters estimated: 5
#> AIC: 376.76
#>
data(boston, package="spData")
gp1 <- spautolm(log(CMEDV) ~ CRIM + ZN + INDUS + CHAS + I(NOX^2)
+ I(RM^2) + AGE + log(DIS) + log(RAD) + TAX + PTRATIO + B + log(LSTAT),
data=boston.c, spdep::nb2listw(boston.soi), family="SMA")
summary(gp1)
#>
#> Call: spautolm(formula = log(CMEDV) ~ CRIM + ZN + INDUS + CHAS + I(NOX^2) +
#> I(RM^2) + AGE + log(DIS) + log(RAD) + TAX + PTRATIO + B +
#> log(LSTAT), data = boston.c, listw = spdep::nb2listw(boston.soi),
#> family = "SMA")
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -0.5847694 -0.0713881 0.0012284 0.0827517 0.6071219
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 4.28501607 0.15367176 27.8842 < 2.2e-16
#> CRIM -0.00718807 0.00106298 -6.7622 1.359e-11
#> ZN 0.00023008 0.00051897 0.4433 0.6575185
#> INDUS 0.00047300 0.00263339 0.1796 0.8574551
#> CHAS1 0.01020698 0.02872047 0.3554 0.7222970
#> I(NOX^2) -0.44885530 0.13675913 -3.2821 0.0010304
#> I(RM^2) 0.00638094 0.00110330 5.7835 7.316e-09
#> AGE -0.00043973 0.00051336 -0.8566 0.3916862
#> log(DIS) -0.15650578 0.03856337 -4.0584 4.941e-05
#> log(RAD) 0.07583760 0.02016468 3.7609 0.0001693
#> TAX -0.00049364 0.00012162 -4.0588 4.933e-05
#> PTRATIO -0.02494959 0.00538791 -4.6307 3.645e-06
#> B 0.00048517 0.00010944 4.4334 9.277e-06
#> log(LSTAT) -0.32961379 0.02353891 -14.0029 < 2.2e-16
#>
#> Lambda: 0.61991 LR test value: 144.28 p-value: < 2.22e-16
#> Numerical Hessian standard error of lambda: 0.04431
#>
#> Log likelihood: 229.1208
#> ML residual variance (sigma squared): 0.02596, (sigma: 0.16112)
#> Number of observations: 506
#> Number of parameters estimated: 16
#> AIC: -426.24
#>
# }