Bootstrapping-based test for local spatial heteroscedasticity
LOSH.mc.RdThe function draws inferences about local spatial heteroscedasticity (LOSH) by means of the randomisation-based Monte-Carlo bootstrap proposed by Xu et al. (2014).
Usage
LOSH.mc(x, listw, a = 2, nsim = 99, zero.policy = attr(listw, "zero.policy"),
na.action = na.fail, spChk = NULL, adjust.n = TRUE, p.adjust.method = "none")Arguments
- x
a numeric vector of the same length as the neighbours list in listw
- listw
a
listwobject created for example bynb2listw- a
the exponent applied to the local residuals; the default value of 2 leads to a measure of heterogeneity in the spatial variance
- nsim
the number of randomisations used in the bootstrap
- zero.policy
default
attr(listw, "zero.policy")as set whenlistwwas created, if attribute not set, use global option value; if TRUE assign zero to the lagged value of zones without neighbours, if FALSE assign NA- na.action
a function (default
na.fail), can also bena.omitorna.exclude- in these cases the weights list will be subsetted to remove NAs in the data. It may be necessary to set zero.policy to TRUE because this subsetting may create no-neighbour observations. Note that only weights lists created without using the glist argument tonb2listwmay be subsetted. Ifna.passis used, zero is substituted for NA values in calculating the spatial lag. (Note that na.exclude will only work properly starting from R 1.9.0, na.omit and na.exclude assign the wrong classes in 1.8.*)- spChk
should the data vector names be checked against the spatial objects for identity integrity, TRUE, or FALSE, default NULL to use
get.spChkOption()- adjust.n
default TRUE, if FALSE the number of observations is not adjusted for no-neighbour observations, if TRUE, the number of observations is adjusted
- p.adjust.method
a character string specifying the probability value adjustment for multiple tests, default "none"; see
p.adjustSP. Note that the number of multiple tests for each region is only taken as the number of neighbours + 1 for each region, rather than the total number of regions.
Details
The test calculates LOSH (see LOSH) and estimates pseudo p-values from a conditional bootstrap. Thereby, the i-th value in each location is held fixed, whereas all other values are permuted nsim times over all other spatial units.
Value
- Hi
LOSH statistic
- E.Hi
expectation of LOSH
- Var.Hi
variance of LOSH
- Z.Hi
the approximately chi-square distributed test statistics
- x_bar_i
local spatially weighted mean values
- ei
residuals about local spatially weighted mean values
- Pr()
p-values for
Hiobtained from a conditional bootstrap distribution
References
Ord, J. K., & Getis, A. 2012. Local spatial heteroscedasticity (LOSH), The Annals of Regional Science, 48 (2), 529–539; Xu, M., Mei, C. L., & Yan, N. 2014. A note on the null distribution of the local spatial heteroscedasticity (LOSH) statistic. The Annals of Regional Science, 52 (3), 697–710.
Author
René Westerholt rene.westerholt@tu-dortmund.de
See also
LOSH, LOSH.mc
Examples
data(columbus, package="spData")
resLOSH_mc <- LOSH.mc(columbus$CRIME, nb2listw(col.gal.nb), 2, 100)
summary(resLOSH_mc)
#> Hi x_bar_i ei Pr()
#> Min. :0.03438 Min. :13.85 Min. :2.982e-02 Min. :0.009901
#> 1st Qu.:0.23838 1st Qu.:24.71 1st Qu.:7.011e+00 1st Qu.:0.079208
#> Median :0.66689 Median :35.90 Median :5.211e+01 Median :0.653465
#> Mean :1.06592 Mean :34.88 Mean :1.519e+02 Mean :0.552233
#> 3rd Qu.:1.59680 3rd Qu.:45.39 3rd Qu.:1.051e+02 3rd Qu.:0.900990
#> Max. :4.68765 Max. :54.91 Max. :2.455e+03 Max. :0.990099
resLOSH_cs <- LOSH.cs(columbus$CRIME, nb2listw(col.gal.nb))
summary(resLOSH_cs)
#> Hi E.Hi Var.Hi Z.Hi
#> Min. :0.03438 Min. :1 Min. :0.4972 Min. :0.04356
#> 1st Qu.:0.23838 1st Qu.:1 1st Qu.:0.9136 1st Qu.:0.35070
#> Median :0.66689 Median :1 Median :1.4342 Median :1.19176
#> Mean :1.06592 Mean :1 Mean :1.4758 Mean :1.89447
#> 3rd Qu.:1.59680 3rd Qu.:1 3rd Qu.:1.9547 3rd Qu.:2.65120
#> Max. :4.68765 Max. :1 Max. :2.9958 Max. :7.25681
#> x_bar_i ei Pr()
#> Min. :13.85 Min. :2.982e-02 Min. :0.0190
#> 1st Qu.:24.71 1st Qu.:7.011e+00 1st Qu.:0.2050
#> Median :35.90 Median :5.211e+01 Median :0.5264
#> Mean :34.88 Mean :1.519e+02 Mean :0.4507
#> 3rd Qu.:45.39 3rd Qu.:1.051e+02 3rd Qu.:0.6529
#> Max. :54.91 Max. :2.455e+03 Max. :0.9540
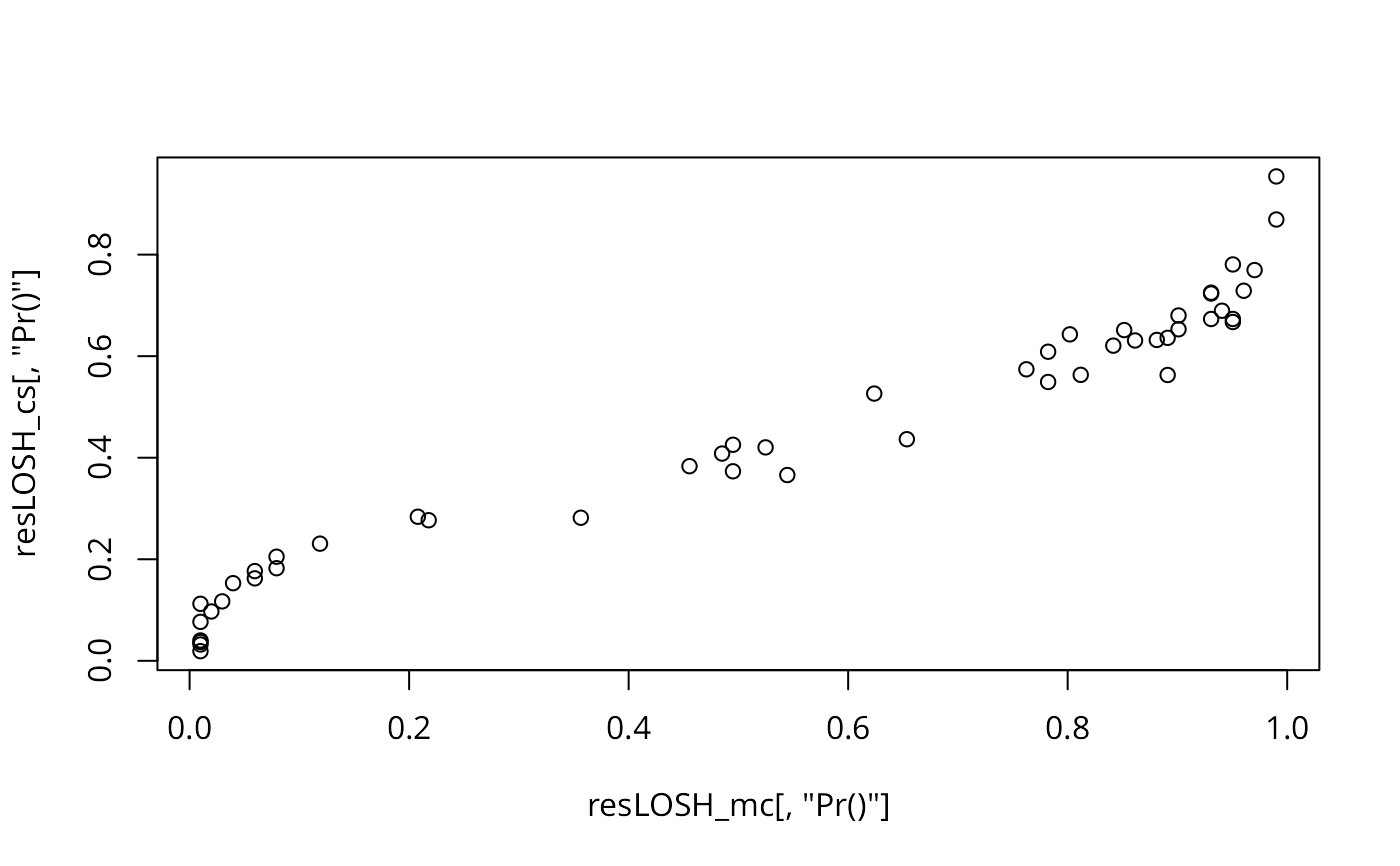
plot(resLOSH_mc[,"Pr()"], resLOSH_cs[,"Pr()"])